How Tplyr Works
When you look at a summary table within a clinical report, you can often break it down into some basic pieces. Consider this output.
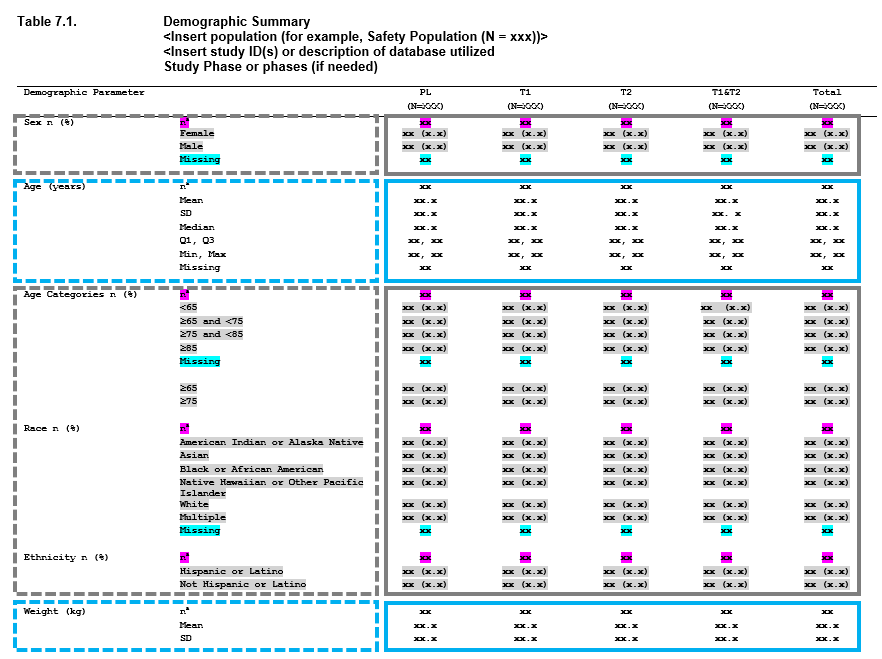
Different variables are being summarized in chunks of the table,
which we refer to as “layers”. Additionally, this table really only
contains a few different types of summaries, which makes many of the
calculations rather redundant. This drives the motivation behind
Tplyr. The containing table is encapsulated within the
tplyr_table() object, and each section, or “layer”, within
the summary table can be broken down into a tplyr_layer()
object.
The tplyr_table() Object
The tplyr_table() object is the conceptual “table” that
contains all of the logic necessary to construct and display the data.
Tplyr tables are made up of one or more layers. Each
layer contains an instruction for a summary to be performed. The
tplyr_table() object contains those layers, and the general
data, metadata, and logic necessary to prepare the data before any
layers are constructed.
When a tplyr_table() is created, it will contain the
following bindings:
-
target- The dataset upon which summaries will be performed -
count_layer_formats- Default formats to be used on count layers in the table -
shift_layer_formats- Default formats to be used on shift layers in the table -
desc_layer_formats- Default formats to be used on descriptive statistics layers in the table -
pop_data- The dataset containing population information. This defaults to thetargetdataset -
cols- A categorical variable in thetargetdataset to present summaries grouped by column (in addition to thetreat_varvariable) -
table_where- Thewhereclause provided, used to subset thetargetdataset -
treat_var- Variable used to distinguish treatment groups in thetargetdataset. -
header_n- Default header N values based ontreat_varand anycolsvariables -
pop_treat_var- Variable used to distinguish treatment groups inpop_datadataset (if different than thetreat_varvariable in thetargetdataset) -
layers- The container for individual layers of atplyr_table() -
treat_grps- Additional treatment groups to be added to the summary (i.e. Total)
The function tplyr_table() allows you a basic interface
to instantiate the object. Modifier functions are available to change
individual parameters catered to your analysis.
t <- tplyr_table(tplyr_adsl, TRT01P, where = SAFFL == "Y")
t
#> *** tplyr_table ***
#> Target (data.frame):
#> Name: tplyr_adsl
#> Rows: 254
#> Columns: 49
#> treat_var variable (quosure)
#> TRT01P
#> header_n: header groups
#> treat_grps groupings (list)
#> Table Columns (cols):
#> where: == SAFFL Y
#> Number of layer(s): 0
#> layer_output: <Table Not Built Yet>The tplyr_layer() Object
Users of Tplyr interface with
tplyr_layer() objects using the
group_<type> family of functions. This family
specifies the type of summary that is to be performed within a layer.
count layers are used to create summary counts of some
discrete variable. shift layers summarize the counts for
different changes in states. Lastly, desc layers create
descriptive statistics.
-
Count Layers
- Count layers allow you to easily create summaries based on counting
distinct or non-distinct occurrences of values within a variable.
Additionally, this layer allows you to create n (%) summaries where
you’re also summarizing the proportion of instances a value occurs
compared to some denominator. Count layers are also capable of producing
counts of nested relationships. For example, if you want to produce
counts of an overall outside group, and then the subgroup counts within
that group, you can simply specify the target variable as
vars(OutsideVariable, InsideVariable). This allows you to do tables like Adverse Events where you want to see the Preferred Terms within Body Systems, all in one layer. Count layers can also distinguish between distinct and non-distinct counts. Using some specified by variable, you can count the unique occurrences of some variable within the specified by grouping, including the target. This allows you to do a summary like unique subjects and their proportion experiencing some adverse event, and the number of total occurrences of that adverse event.
- Count layers allow you to easily create summaries based on counting
distinct or non-distinct occurrences of values within a variable.
Additionally, this layer allows you to create n (%) summaries where
you’re also summarizing the proportion of instances a value occurs
compared to some denominator. Count layers are also capable of producing
counts of nested relationships. For example, if you want to produce
counts of an overall outside group, and then the subgroup counts within
that group, you can simply specify the target variable as
-
Descriptive Statistics Layers
- Descriptive statistics layers perform summaries on continuous
variables. There are a number of summaries built into
Tplyr already that you can perform, including n, mean,
median, standard deviation, variance, min, max, interquartile range, Q1,
Q3, and missing value counts. From these available summaries, the
default presentation of a descriptive statistics layer will output ‘n’,
‘Mean (SD)’, ‘Median’, ‘Q1, Q3’, ‘Min, Max’, and ‘Missing’. You can
change these summaries using
set_format_strings(), and you can also add your own summaries usingset_custom_summaries(). This allows you to easily implement any additional summary statistics you want presented.
- Descriptive statistics layers perform summaries on continuous
variables. There are a number of summaries built into
Tplyr already that you can perform, including n, mean,
median, standard deviation, variance, min, max, interquartile range, Q1,
Q3, and missing value counts. From these available summaries, the
default presentation of a descriptive statistics layer will output ‘n’,
‘Mean (SD)’, ‘Median’, ‘Q1, Q3’, ‘Min, Max’, and ‘Missing’. You can
change these summaries using
-
Shift Layers
- Shift layers are largely an abstraction of the count layer - and in fact, we re-use a lot of the same code to process these layers. In many shift tables, the “from” state is presented as rows in the table, and the “to” state is presented as columns. This clearly lays out how many subjects changed state between a baseline and some point in time. Shift layers give you an intuitive API to break these out, using a very similar interface as the other layers. There are also a number of modifier functions available to control nuanced aspects, such as how denominators should be applied.
cnt <- group_count(t, AGEGR1)
cnt
#> *** count_layer ***
#> Self: count_layer < 0x564fd85f05d8 >
#> Parent: tplyr_table < 0x564fd82946c8 >
#> target_var:
#> AGEGR1
#> by:
#> where: TRUE
#> Layer(s): 0
dsc <- group_desc(t, AGE)
dsc
#> *** desc_layer ***
#> Self: desc_layer < 0x564fd8754748 >
#> Parent: tplyr_table < 0x564fd82946c8 >
#> target_var:
#> AGE
#> by:
#> where: TRUE
#> Layer(s): 0
shf <- group_shift(t, vars(row=COMP8FL, column=COMP24FL))
shf
#> *** shift_layer ***
#> Self: shift_layer < 0x564fd8858920 >
#> Parent: tplyr_table < 0x564fd82946c8 >
#> target_var:
#> COMP8FL
#> COMP24FL
#> by:
#> where: TRUE
#> Layer(s): 0Adding Layers to a Table
Everyone has their own style of coding - so we’ve tried to be
flexible to an extent. Overall, Tplyr is built around
tidy syntax, so all of our object construction supports piping with magrittr
(i.e. %>%).
There are two ways to add layers to a tplyr_table():
add_layer() and add_layers(). The difference
is that add_layer() allows you to construct the layer
within the call to add_layer(), whereas with
add_layers() you can attach multiple layers that have
already been constructed upfront:
t <- tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories n (%)")
)Within add_layer(), the syntax to constructing the count
layer for Age Categories was written on the fly.
add_layer() is special in that it also allows you to use
piping to use modifier functions on the layer being constructed
t <- tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories n (%)") %>%
set_format_strings(f_str("xx (xx.x%)", n, pct)) %>%
add_total_row()
)add_layers(), on the other hand, lets you isolate the
code to construct a particular layer if you wanted to separate things
out more. Some might find this cleaner to work with if you have a large
number of layers being constructed.
t <- tplyr_table(tplyr_adsl, TRT01P)
l1 <- group_count(t, AGEGR1, by = "Age categories n (%)")
l2 <- group_desc(t, AGE, by = "Age (years)")
t <- add_layers(t, l1, l2)Notice that when you construct the layers separately, you need to
specify the table to which they belong. add_layer() does
this automatically. tplyr_table() and
tplyr_layer() objects are built on environments, and the
parent/child relationships are very important. This is why, even though
the layer knows who its table parent is, the layers still need to be
attached to the table (as the table doesn’t know who its children are).
Advanced R does
a very good job at explaining what environments in R are, their
benefits, and how to use them.
A Note Before We Go Deeper
Notice that when you construct a tplyr_table() or a
tplyr_layer() that what displays is a summary of
information about the table or layer? That’s because when you create
these objects - it constructs the metadata, but does not process the
actual data. This allows you to construct and make sure the pieces of
your table fit together before you do the data processing - and it gives
you a container to hold all of this metadata, and use it later if
necessary.
To generate the data from a tplyr_table() object, you
use the function build():
t <- tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories n (%)")
)
t %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age categories n (%) | <65 | 14 ( 16.3%) | 11 ( 13.1%) | 8 ( 9.5%) | 1 | 1 | 1 |
| Age categories n (%) | >80 | 30 ( 34.9%) | 18 ( 21.4%) | 29 ( 34.5%) | 1 | 1 | 2 |
| Age categories n (%) | 65-80 | 42 ( 48.8%) | 55 ( 65.5%) | 47 ( 56.0%) | 1 | 1 | 3 |
But there’s more you can get from Tplyr. It’s great
to have the formatted numbers, but what about the numeric data behind
the scenes? Maybe a number looks suspicious and you need to investigate
how you got that number. What if you want to calculate your own
statistics based off of the counts? You can get that information as well
using get_numeric_data(). This returns the numeric data
from each layer as a list of data frames:
get_numeric_data(t) %>%
head() %>%
kable()
|
By storing pertinent information, you can get more out of a Tplyr object than processed data for display. And by specifying when you want to get data out of Tplyr, we can save you from repeatedly processing data while your constructing your outputs - which is particularly useful when that computation starts taking time.
Constructing Layers
The bulk of Tplyr coding comes from constructing your layers and specifying the work you want to be done. Before we get into this, it’s important to discuss how Tplyr handles string formatting.
String Formatting in Tplyr
String formatting in Tplyr is controlled by an
object called an f_str(), which is also the name of
function you use to create these formats. To set these format strings
into a tplyr_layer(), you use the function
set_format_strings(), and this usage varies slightly
between layer types (which is covered in other vignettes).
So - why is this object necessary. Consider this example:
t <- tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_desc(AGE, by = "Age (years)") %>%
set_format_strings(
'n' = f_str('xx', n),
'Mean (SD)' = f_str('xx.xx (xx.xxx)', mean, sd)
)
)
t %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age (years) | n | 86 | 84 | 84 | 1 | 1 | 1 |
| Age (years) | Mean (SD) | 75.21 ( 8.590) | 74.38 ( 7.886) | 75.67 ( 8.286) | 1 | 1 | 2 |
In a perfect world, the f_str() calls wouldn’t be
necessary - but in reality they allow us to infer a great deal of
information from very few user inputs. In the calls that you see
above:
- The row labels in the
row_label2column are taken from the left side of each=inset_format_strings() - The string formats, including integer length and decimal precision,
and exact presentation formatting are taken from the strings within the
first parameter of each
f_str()call - The second and greater parameters within each
f_str()call determine the descriptive statistic summaries that will be performed. This is connected to a number of default summaries available within Tplyr, but you can also create your own summaries (covered in other vignettes). The default summaries that are built in include:-
n= Number of observations -
mean= Mean -
sd= Standard Deviation -
var= Variance -
iqr= Inter Quartile Range -
q1= 1st quartile -
q3= 3rd quartile -
min= Minimum value -
max= Maximum value -
missing= Count of NA values
-
- When two summaries are placed on the same
f_str()call, then those two summaries are formatted into the same string. This allows you to do a “Mean (SD)” type format where both numbers appear.
This simple user input controls a significant amount of work in the
back end of the data processing, and the f_str() object
allows that metadata to be collected.
f_str() objects are also used with count layers as well
to control the data presentation. Instead of specifying the summaries
performed, you use n, pct,
distinct_n, and distinct_pct for your
parameters and specify how you would like the values displayed. Using
distinct_n and distinct_pct should be combined
with specifying a distinct_by() variable using
set_distinct_by().
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories") %>%
set_format_strings(f_str('xx (xx.x)',n,pct))
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age categories | <65 | 14 (16.3) | 11 (13.1) | 8 ( 9.5) | 1 | 1 | 1 |
| Age categories | >80 | 30 (34.9) | 18 (21.4) | 29 (34.5) | 1 | 1 | 2 |
| Age categories | 65-80 | 42 (48.8) | 55 (65.5) | 47 (56.0) | 1 | 1 | 3 |
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories") %>%
set_format_strings(f_str('xx',n))
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age categories | <65 | 14 | 11 | 8 | 1 | 1 | 1 |
| Age categories | >80 | 30 | 18 | 29 | 1 | 1 | 2 |
| Age categories | 65-80 | 42 | 55 | 47 | 1 | 1 | 3 |
Really - format strings allow you to present your data however you like.
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories") %>%
set_format_strings(f_str('xx (•◡•) xx.x%',n,pct))
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age categories | <65 | 14 (•◡•) 16.3% | 11 (•◡•) 13.1% | 8 (•◡•) 9.5% | 1 | 1 | 1 |
| Age categories | >80 | 30 (•◡•) 34.9% | 18 (•◡•) 21.4% | 29 (•◡•) 34.5% | 1 | 1 | 2 |
| Age categories | 65-80 | 42 (•◡•) 48.8% | 55 (•◡•) 65.5% | 47 (•◡•) 56.0% | 1 | 1 | 3 |
But should you? Probably not.
Layer Types
Descriptive Statistic Layers
As covered under string formatting, set_format_strings()
controls a great deal of what happens within a descriptive statistics
layer. Note that there are some built in defaults to what’s output:
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_desc(AGE, by = "Age (years)")
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age (years) | n | 86 | 84 | 84 | 1 | 1 | 1 |
| Age (years) | Mean (SD) | 75.2 ( 8.59) | 74.4 ( 7.89) | 75.7 ( 8.29) | 1 | 1 | 2 |
| Age (years) | Median | 76.0 | 76.0 | 77.5 | 1 | 1 | 3 |
| Age (years) | Q1, Q3 | 69.2, 81.8 | 70.8, 80.0 | 71.0, 82.0 | 1 | 1 | 4 |
| Age (years) | Min, Max | 52, 89 | 56, 88 | 51, 88 | 1 | 1 | 5 |
| Age (years) | Missing | 0 | 0 | 0 | 1 | 1 | 6 |
To override these defaults, just specify the summaries that you want
to be performed using set_format_strings() as described
above. But what if Tplyr doesn’t have a built in
function to do the summary statistic that you want to see? Well - you
can make your own! This is where set_custom_summaries()
comes into play. Let’s say you want to derive a geometric mean.
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_desc(AGE, by = "Sepal Length") %>%
set_custom_summaries(
geometric_mean = exp(sum(log(.var[.var > 0]), na.rm=TRUE) / length(.var))
) %>%
set_format_strings(
'Geometric Mean (SD)' = f_str('xx.xx (xx.xxx)', geometric_mean, sd)
)
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Sepal Length | Geometric Mean (SD) | 74.70 ( 8.590) | 73.94 ( 7.886) | 75.18 ( 8.286) | 1 | 1 | 1 |
In set_custom_summaries(), first you name the summary
being performed. This is important - that name is what you use in the
f_str() call to incorporate it into a format. Next, you
program or call the function desired. What happens in the background is
that this is used in a call to dplyr::summarize() - so use
similar syntax. Use the variable name .var in your custom
summary function. This is necessary because it allows a generic variable
name to be used when multiple target variables are specified - and
therefore the function can be applied to both target variables.
Sometimes there’s a need to present multiple variables summarized side by side. Tplyr allows you to do this as well.
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_desc(vars(AGE, AVGDD), by = "Age and Avg. Daily Dose")
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | var2_Placebo | var2_Xanomeline High Dose | var2_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|---|---|---|
| Age and Avg. Daily Dose | n | 86 | 84 | 84 | 86 | 84 | 84 | 1 | 1 | 1 |
| Age and Avg. Daily Dose | Mean (SD) | 75.2 ( 8.59) | 74.4 ( 7.89) | 75.7 ( 8.29) | 0.0 ( 0.00) | 71.6 ( 8.11) | 54.0 ( 0.00) | 1 | 1 | 2 |
| Age and Avg. Daily Dose | Median | 76.0 | 76.0 | 77.5 | 0.0 | 75.1 | 54.0 | 1 | 1 | 3 |
| Age and Avg. Daily Dose | Q1, Q3 | 69.2, 81.8 | 70.8, 80.0 | 71.0, 82.0 | 0.0, 0.0 | 70.2, 76.9 | 54.0, 54.0 | 1 | 1 | 4 |
| Age and Avg. Daily Dose | Min, Max | 52, 89 | 56, 88 | 51, 88 | 0, 0 | 54, 79 | 54, 54 | 1 | 1 | 5 |
| Age and Avg. Daily Dose | Missing | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 6 |
Tplyr summarizes both variables and merges them
together. This makes creating tables where you need to compare BASE,
AVAL, and CHG next to each other nice and simple. Note the use of
dplyr::vars() - in any situation where you’d like to use
multiple variable names in a parameter, use dplyr::vars()
to specify the variables. You can use text strings in the calls to
dplyr::vars() as well.
Count Layers
Count layers generally allow you to create “n” and “n (%)” count type
summaries. There are a few extra features here as well. Let’s say that
you want a total row within your counts. This can be done with
add_total_row():
tplyr_table(tplyr_adsl, TRT01P) %>%
add_layer(
group_count(AGEGR1, by = "Age categories") %>%
add_total_row()
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| Age categories | <65 | 14 ( 16.3%) | 11 ( 13.1%) | 8 ( 9.5%) | 1 | 1 | 1 |
| Age categories | >80 | 30 ( 34.9%) | 18 ( 21.4%) | 29 ( 34.5%) | 1 | 1 | 2 |
| Age categories | 65-80 | 42 ( 48.8%) | 55 ( 65.5%) | 47 ( 56.0%) | 1 | 1 | 3 |
| Age categories | Total | 86 (100.0%) | 84 (100.0%) | 84 (100.0%) | 1 | 1 | 4 |
Sometimes it’s also necessary to count summaries based on distinct
values. Tplyr allows you to do this as well with
set_distinct_by():
tplyr_table(tplyr_adae, TRTA) %>%
add_layer(
group_count('Subjects with at least one adverse event') %>%
set_distinct_by(USUBJID) %>%
set_format_strings(f_str('xx', n))
) %>%
build() %>%
kable()| row_label1 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 |
|---|---|---|---|---|---|
| Subjects with at least one adverse event | 47 | 111 | 118 | 1 | NA |
There’s another trick going on here - to create a summary with row
label text like you see above, text strings can be used as the target
variables. Here, we use this in combination with
set_distinct_by() to count distinct subjects.
Adverse event tables often call for counting AEs of something like a body system and counting actual events within that body system. Tplyr has means of making this simple for the user as well.
tplyr_table(tplyr_adae, TRTA) %>%
add_layer(
group_count(vars(AEBODSYS, AEDECOD))
) %>%
build() %>%
head() %>%
kable()| row_label1 | row_label2 | var1_Placebo | var1_Xanomeline High Dose | var1_Xanomeline Low Dose | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | SKIN AND SUBCUTANEOUS TISSUE DISORDERS | 47 (100.0%) | 111 (100.0%) | 118 (100.0%) | 1 | 1 | Inf |
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | ACTINIC KERATOSIS | 0 ( 0.0%) | 1 ( 0.9%) | 0 ( 0.0%) | 1 | 1 | 1 |
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | ALOPECIA | 1 ( 2.1%) | 0 ( 0.0%) | 0 ( 0.0%) | 1 | 1 | 2 |
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | BLISTER | 0 ( 0.0%) | 2 ( 1.8%) | 8 ( 6.8%) | 1 | 1 | 3 |
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | COLD SWEAT | 3 ( 6.4%) | 0 ( 0.0%) | 0 ( 0.0%) | 1 | 1 | 4 |
| SKIN AND SUBCUTANEOUS TISSUE DISORDERS | DERMATITIS ATOPIC | 1 ( 2.1%) | 0 ( 0.0%) | 0 ( 0.0%) | 1 | 1 | 5 |
Here we again use dplyr::vars() to specify multiple
target variables. When used in a count layer, Tplyr
knows automatically that the first variable is a grouping variable for
the second variable, and counts shall be produced for both then merged
together.
Shift Layers
Lastly, let’s talk about shift layers. A common example of this would be looking at a subject’s lab levels at baseline versus some designated evaluation point. This would tell us, for example, how many subjects were high at baseline for a lab test vs. after an intervention has been introduced. The shift layer in Tplyr is intended for creating shift tables that show these data as a matrix, where one state will be presented in rows and the other in columns. Let’s look at an example.
# Tplyr can use factor orders to dummy values and order presentation
tplyr_adlb$ANRIND <- factor(tplyr_adlb$ANRIND, c("L", "N", "H"))
tplyr_adlb$BNRIND <- factor(tplyr_adlb$BNRIND, c("L", "N", "H"))
tplyr_table(tplyr_adlb, TRTA, where = PARAMCD == "CK") %>%
add_layer(
group_shift(vars(row=BNRIND, column=ANRIND), by=PARAM) %>%
set_format_strings(f_str("xx (xxx%)", n, pct))
) %>%
build() %>%
kable()| row_label1 | row_label2 | var1_Placebo_L | var1_Placebo_N | var1_Placebo_H | var1_Xanomeline High Dose_L | var1_Xanomeline High Dose_N | var1_Xanomeline High Dose_H | var1_Xanomeline Low Dose_L | var1_Xanomeline Low Dose_N | var1_Xanomeline Low Dose_H | ord_layer_index | ord_layer_1 | ord_layer_2 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Creatine Kinase (U/L) | L | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 1 | 35 | 1 |
| Creatine Kinase (U/L) | N | 0 ( 0%) | 27 ( 87%) | 4 ( 13%) | 0 ( 0%) | 17 ( 85%) | 2 ( 10%) | 0 ( 0%) | 14 ( 93%) | 1 ( 7%) | 1 | 35 | 2 |
| Creatine Kinase (U/L) | H | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 1 ( 5%) | 0 ( 0%) | 0 ( 0%) | 0 ( 0%) | 1 | 35 | 3 |
The underlying process of shift tables is the same as count layers -
we’re counting the number of occurrences of something by a set of
grouping variables. This differs in that Tplyr uses the
group_shift() API to use the same basic interface as other
tables, but translate your target variables into the row variable and
the column variable. Furthermore, there is some enhanced control over
how denominators should behave that is necessary for a shift layer.
Where to go from here?
There’s quite a bit more to learn! And we’ve prepared a number of other vignettes to help you get what you need out of Tplyr.
- Learn more about table level settings in
vignette("table") - Learn more about descriptive statistics layers in
vignette("desc") - Learn more about count and shift layers in
vignette("count") - Learn more about shift layers in
vignette("shift") - Learn more about calculating risk differences in
vignette("riskdiff") - Learn more about sorting Tplyr tables in
vignette("sort") - Learn more about using Tplyr options in
vignette("options") - And finally, learn more about producing and outputting styled tables
using Tplyr in
vignette("styled-table")
References
In building Tplyr, we needed some additional resources in addition to our personal experience to help guide design. PHUSE has done some great work to create guidance for standard outputs with collaboration between multiple pharmaceutical companies and the FDA. You can find some of the resource that we referenced below.
Analysis and Displays Associated with Adverse Events
Analyses and Displays Associated with Demographics, Disposition, and Medications
Analyses and Displays Associated with Measures of Central Tendency
